Group Schuijers

Areas of expertise uitklapper, klik om te openen
Dr. Jurian Schuijers is group leader at the department of Molecular Cancer Research within the Center of Molecular Medicine at the UMC Utrecht. He obtained his PhD under supervision of Prof. Dr. Hans Clevers (Hubrecht Institute ), where he gained an interest in signaling-driven transcriptional regulation in intestinal stem cells. He studied the interactions between β-catenin and TCF, the role of OLFM4 in intestinal stem cells and elucidated a transcriptional network that acts as a master switch to impart stem cell properties.
During his postdoc in the lab of Prof. Dr. Richard Young (Whitehead institute/MIT) he continued studying signaling-driven transcription and uncovered how super-enhancers, clusters of enhancers that accumulate a disproportionate amount of transcriptional co-factors, serve as signal integration platforms for key cell identity genes. He studied 3D-DNA-architecture and how this affects gene regulation. He showed that enhancer-promoter looping can be dependent on CTCF-occupied enhancer-docking sites that can be perturbed to inhibit gene expression. He studied how components of the transcriptional machinery and transcription factors form biomolecular condensates and showed that signaling factors can concentrated in these condensates to achieve context dependent interpretation of signaling.
Research program uitklapper, klik om te openen
Transcriptional control is one of the major determining factors for correct development and cellular differentiation. As such, failure to achieve appropriate transcriptional control lies at the heart of the majority of diseases. By studying the mechanisms by which transcriptional control is achieved we aim to gain insights in these processes with as ultimate goal therapeutic intervention.
Signaling condensates
Many cellular processes are organized in localized membraneless compartments called biomolecular condensates. We study the nuclear, cytoplasmic and membrane-associated condensates involved in key signaling pathways such as Wnt, Tgf-b and JAK/STAT. We have developed live-cell imaging techniques to uncover the dynamics and function of these condensates.
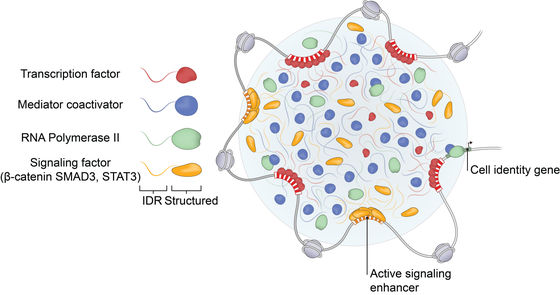
3D-DNA-Architecture
During interphase the genome is packed into the nucleus in a highly organized manner. This 3D-architecture of the DNA plays an important role in transcriptional regulation. Using 4C and Hi-C techniques in combination with FISH based microscopy we study how 3D-DNA structures and nuclear condensates interact.
Molecular features underlying condensate formation
The capacity to undergo liquid-liquid phase separation (LLPS) is likely to be the driving mechanism for the formation of biomolecular condensates. In order to understand the behavior of signaling condensates we study which molecular features of the terminal signaling factors are important for their propensity to undergo LLPS. Using in vitro droplet formation assays and in vitro perturbation assays we analyze the importance of amino acid side chains, phosphorylation, and backbone flexibility for condensate formation.
Key Publications uitklapper, klik om te openen
- Zamudio AV, Dall’Agnese A, Henninger JE, Manteiga JC, Afeyan LK, Hannet N, Coffey EL, Li CH, Oksuz O, Sabari BR, Boija A, Klein IA, Hawken SW, Spillie JH, Decker TM, Cisse II, Abraham BJ, Lee TJ, Taatjes DJ, Schuijers J*§, Young RA*§. Mediator condensates localize signaling factors to key cell identity genes. Mol. Cell. 2019 Dec 5. Dec 05; 76(5) 753-766.e6
- Boija A, Klein IA, Sabari BR, Dall'Agnese A, Coffey EL, Zamudio AV, Li CH, Shrinivas K, Manteiga JC, Hannett NM, Abraham BJ, Afeyan LK, Guo YE, Rimel JK, Fant CB, Schuijers J, Lee TI, Taatjes DJ, Young RA. Transcription Factors Activate Genes through the Phase-Separation Capacity of Their Activation Domains. Cell. 2018 Dec 13;175(7):1842-1855.e16.
- Sabari BR, Dall'Agnese A, Boija A, Klein IA, Coffey EL, Shrinivas K, Abraham BJ, Hannett NM, Zamudio AV, Manteiga JC, Li CH, Guo YE, Day DS, Schuijers J, Vasile E, Malik S, Hnisz D, Lee TI, Cisse II, Roeder RG, Sharp PA, Chakraborty AK, Young RA. Coactivator condensation at super-enhancers links phase separation and gene control. Science. 2018 Jul 27;361(6400).
- Schuijers J*, Manteiga JC*, Weintraub AS, Day DS, Zamudio AV, Hnisz D, Lee TI, Young RA. Transcriptional Dysregulation of MYC Reveals Common Enhancer-Docking Mechanism. Cell Rep. 2018 Apr 10;23(2):349-360.
- Hnisz D*, Schuijers J*, Lin CY, Weintraub AS, Abraham BJ, Lee TI, Bradner JE, Young RA. Convergence of developmental and oncogenic signaling pathways at transcriptional super-enhancers. Mol Cell. 2015 Apr 16;58(2):362-70
- Schuijers J, Junker JP, Mokry M, Hatzis P, Koo BK, Sasselli V, van der Flier LG, Cuppen E, van Oudenaarden A, Clevers H. Ascl2 acts as an R-spondin/Wnt-responsive switch to control stemness in intestinal crypts. Cell Stem Cell. 2015 Feb 5;16(2):158-70
Full list of publications here
Lab website
Group members uitklapper, klik om te openen
- Now Hiring! – PhD student
- Tianhshu Gui - PhD student shared with Burgering lab
- Jasper Heuer - student
Visiting address:
Stratenum 2.131
Universiteitsweg 100
3584 CG Utrecht
The Netherlands
Secretariat:
Marianne van der Heiden
a.m.vanderheiden@umcutrecht.nl
+31 (0)88 75 68989
